Immunology Ontologies: Difference between revisions
mNo edit summary |
|||
| (2 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
== | == ImmPort == | ||
This page was initially created to support the ontology work of the Immunology Database and Analysis Portal ([https://immport.niaid.nih.gov/ ImmPort]) portal within the framework of the Bioinformatics Integration Support Contract ([https://www.fbo.gov/index?s=opportunity&mode=form&id=67aa839f87c97f95f7531ba9d86f67b5&tab=core&_cview=1 BISC]), funded by the NIH’s National Institute of Allergy and Infectious Diseases (NIAID). ImmPort is designed to enable scientists to easily access and exchange interoperable complex data sets to accelerate scientific discovery. It provides advanced information technology support in the production, analysis, archiving, and exchange of scientific data for the diverse community of life science researchers supported by NIAID. A summary of work so far is provided here: | |||
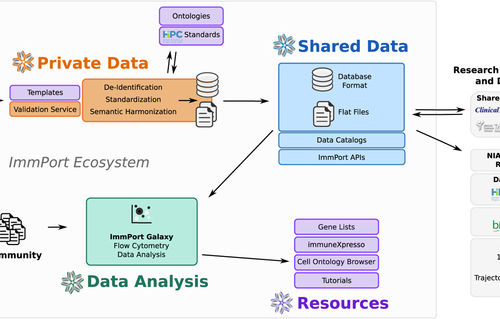
'''[https://www.nature.com/articles/sdata201815 ImmPort, toward repurposing of open access immunological assay data for translational and clinical research'''] | |||
[[File:sdata201815-f1.jpg]] | [[File:sdata201815-f1.jpg]] | ||
See also '''[http://ncorwiki.buffalo.edu/index.php/ImmPort_Ontology_Conference ImmPort Ontology Conference]''' | |||
== Background == | |||
The huge plasticity of the immune response, the strong dependence upon the context in which it occurs and on physiological and pathological conditions, and the wide range of responses elicited, make immunology an especially challenging domain for data integration. | The huge plasticity of the immune response, the strong dependence upon the context in which it occurs and on physiological and pathological conditions, and the wide range of responses elicited, make immunology an especially challenging domain for data integration. | ||
Latest revision as of 18:50, 4 March 2018
ImmPort
This page was initially created to support the ontology work of the Immunology Database and Analysis Portal (ImmPort) portal within the framework of the Bioinformatics Integration Support Contract (BISC), funded by the NIH’s National Institute of Allergy and Infectious Diseases (NIAID). ImmPort is designed to enable scientists to easily access and exchange interoperable complex data sets to accelerate scientific discovery. It provides advanced information technology support in the production, analysis, archiving, and exchange of scientific data for the diverse community of life science researchers supported by NIAID. A summary of work so far is provided here:
See also ImmPort Ontology Conference
Background
The huge plasticity of the immune response, the strong dependence upon the context in which it occurs and on physiological and pathological conditions, and the wide range of responses elicited, make immunology an especially challenging domain for data integration.
The ontologies documented below are designed to enhance the degree to which immunology research results can be expressed in a consistent fashion across communities and disciplines, thus advancing not only the analysis and integration of data but also its discoverability by scientists who were not involved in its creation. In this way, they will contribute to the realization of the goals of ImmPort to accelerate a more collaborative and coordinated research environment and create an integrated database. that broadens the usefulness of scientific data and advances hypothesis-driven and hypothesis-generating research, as well as the development of optimal methods for data collection, storage, exchange and interoperability.
The team of University at Buffalo ontologists with primary responsibility for this work includes:
- Barry Smith, Professor of Philosophy, Neurology and Computer Science and Director of the National Center for Ontological Research.
- Alexander Diehl, UB Department of Neurology
- Alan Ruttenberg, UB Department of Oral Biology.
For further background see: http://www.buffalo.edu/news/releases/2009/01/9857.html
Ontologies
Recommended
Chemical Entities of Biological Interest (CHEBI) (Paper)
The Protein Ontology (PRO) Immunology Content in the Protein Ontology (Paper)
The Gene Ontology (GO)
The Cell Ontology (CL)
The Immune Epitope Ontology (ONTIE) (paper)
Infectious Disease Ontology (IDO)
Ontology of Vaccine Adverse Events (Paper)
Ontology for Biomedical Investigations (OBI) (Paper)
Ontology for General Medical Science (OGMS)
Under development
Ontology Consortia
OBO (Open Biological and Biomedical Ontologies) Foundry
Clinical Trial Ontology / CDISC
Ravi Shankar, Clinical Trial Ontology, March 13, 2014
CDISC Phuse paper on LabTests, 2010
Clinical Trial Ontology Meeting, 2007
Portals
Immunology Database and Analysis (ImmPort) Portal
NCBO Bioportal (ontology repository)
Ontobee (ontology visualization portal)
HIPC: Human Immunology Project Consortium
Program for Research on Immune Modeling and Experimentation PRIME
Immune Epitope Database and Analysis Resource
International ImMunoGeneTics Information System
ImmPort Antibody Registry
Alex Diehl, The ImmPort Antibody Registry and Ontology
Cytokines
Barry Smith, The OBO Foundry approach to ontologies and standards with special reference to cytokines
Laboratory Information Systems
Open Source
- LabKey
- OpenClinica
- caLIMS (from NCI)
- Labmatica
- Bika
Proprietary
Sample process focused:
Publications
Maecker, H., McCoy, J.P. & Nussenblatt, R. Standardizing immunophenotyping for the human immunology project. Nature reviews Immunology 12, 191-200 (2012).
Smith B, Ashburner M, Rosse C, Bard J, Bug W, Ceusters W, Goldberg LJ, Eilbeck K, Ireland A, Mungall CJ; OBI Consortium, Leontis N, Rocca-Serra P, Ruttenberg A, Sansone SA, Scheuermann RH, Shah N, Whetzel PL, Lewis S. , The OBO Foundry: Coordinated Evolution of Ontologies to Support Biomedical Data Integration, Nature Biotechnology, 25 (11), November 2007, 1251-1255.
Cowell LG, Smith B, Infectious Disease Ontology, in Vitali Sintchenko, Infectious Disease Informatics, New York: Springer, December 2009, 373-395.
Masci AM, Arighi CN, Diehl AD, Lieberman AE, Mungall C, Scheuermann RH, Smith B, Cowell LG, An improved ontological representation of dendritic cells as a paradigm for all cell types, BMC Bioinformatics, February 2009, 10:70.
Diehl AD, Augustine AD, Blake JA, Cowell LG, et al. Hematopoietic cell types: prototype for a revised cell ontology. J Biomed Inform. 2011; 44(1).
Meehan TF, Masci AM, Abdulla A, Cowell LG, et al. Logical development of the cell ontology. BMC Bioinformatics. 2011; 12.
Goldfain A, Smith B, Cowell LG, Towards an Ontological Representation of Resistance: The Case of MRSA, Journal of Biomedical Informatics, 2011 (Feb.), 44:1, 35-41.
Scheuermann RH, Ceusters W, Smith B, Toward an Ontological Treatment of Disease and Diagnosis, Proceedings of the 2009 AMIA Summit on Translational Bioinformatics, 2009, 116-120.
Goldfain A, Smith B, Cowell LG, Dispositions and the Infectious Disease Ontology, in Antony Galton and Riichiro Mizoguchi (eds.), Formal Ontology in Information Systems. Proceedings of the Sixth International Conference (FOIS 2010), Amsterdam: IOS Press, 2010, 400-413.
Goldfain A, Smith B, Cowell LG, Constructing a Lattice of Infectious Disease Ontologies from a Staphylococcus aureus Isolate Repository”, Proceeedings of the Third International Conference on Biomedical Ontology, Graz, July 23-25, 2012 (CEUR, vol. 897).
Cox AP, Jensen M, Duncan W, Weinstock-Guttman B, Szigeti K, Ruttenberg A, Smith B, Diehl AD, Ontologies for the Study of Neurological Disease, Towards an Ontology of Mental Functioning (ICBO Workshop), Third International Conference on Biomedical Ontology, Graz, July 22, 2012.
Yu AC, Smith B, Schwartz S, [http://ontology.buffalo.edu/medo/ACAAI-2012.pdf Formal and Computable Representations of Allergic Diseases in the Electronic Health Record: An Approach Based on the Ontology of General Medical Science, 2012 Annual Meeting of the American College of Allergy, Asthma & Immunology (ACAAI), November 8-13, 2012, Anaheim, California (Poster).
Events
June 11-13, 2012
April 8, 2013
- 1st ImmPort Ontology Group Web Conference:
- 4:00pm Lindsay Cowell (UT Southwestern): The VDJ Server
- 5:00pm General discussion of CL and CyTOF
April 22, 2013
- 2nd ImmPort Ontology Group Web Conference
- 4:00pm Nikolay Samusik (Stanford): Charting the reference map of the immune system using CyTOF data
April 29, 2013
- 3rd Immport Ontology Group Web Conference
- 4:00pm Yannick Pouliot (Stanford): Peak Probability Contrasts (PPC)
June 11, 2013
- Lecture and practical session on Immunology Ontology
- Summer School for Quantitative Systems Immunology, Boston, MA.
June 24, 2013
August 1, 2013
- How to make ImmPort data fit for secondary use, Barry Smith
- Immport Science Group
September 4-5, 2013
- ImmPort Ontology Conference, Stanford University.
October 3, 2013 Ontology Update: Antibodies, Proteins Cells, Alexender Diehl
- ImmPort Science Group
Oct 9-10, 2013
- Training and Strategy Workshop for ImmPort Data Submitters, Rho Federal Systems Division, Chapel Hill, NC, October 9-10
January 21, 2014
- Clinical Trial Data Wants to be Free
- Presentation by Barry Smith, University at Buffalo Clinical and Research Ethics Seminar
February 27, 2014
- Enhancing the Quality of ImmPort Data, Immport Science Group
March 13, 2014
- 5th ImmPort Ontology Group Web Conference
- Presentation by Ravi Shankar on the Clinical Trial Ontology
March 18, 2014
March 20, 2014
- Gene Expression and Cell Identity, Alexender Diehl
- ImmPort Science Group
April 3, 2014
- Ontology and the Future of Laboratory Information, David Parrish (sampleminded.com)
- Buffalo CTRC
Powerpoint Presentations
Barry Smith, Strategies to Enhance Discoverability of Clinical Trial Data